58.4.1. 1-dimensional P-system#
58.4.1.1. Useful classes for numerical methods#
58.4.1.1.1. Basic libraries#
Show code cell source
import numpy as np
import matplotlib.pyplot as plt
58.4.1.1.2. Parent class#
\(\texttt{HyperbolicSystem1d()}\) class with common functions (or just their templates) required to implement a finite volume method solver.
Show code cell source
class HyperbolicSystem1d():
"""
Analytical quantities of a Hyperbolic linear system in a 1-dimensional domain
"""
def __init(self, **params):
pass
def flux(self,):
""" Analytical flux, f """
pass
def A(self,):
""" Advection matrix, A """
pass
def s(self,):
""" Eigenvalues of matrix, s """
pass
def R(self,):
""" Matrix of right eigenvectors, R """
pass
def L(self,):
""" Matrix of left eigenvectors, L=inv(R) """
pass
def spectrum(self,):
""" Spectrum of matrix A() """
return self.s(), self.R(), self.L()
def absA(self, u):
""" A = R * S * L, |A| = R * |S| * L """
return self.R(u) @ np.diag(np.abs(self.s(u))) @ self.L(u)
def roe_intermediate_state(self,):
""" Roe intermediate state for Roe linearization """
pass
58.4.1.1.3. P-system class#
Show code cell source
class Psys1d(HyperbolicSystem1d):
"""
Analytical quantities of a Hyperbolic linear system in a 1-dimensional domain
Physical quantities:
- conservative: (rho, m)
- physical/convective (rho, u)
Parameters:
- a: speed of sound
"""
def __init__(self, a, name=""):
""" """
self.a = a
self.a2 = self.a**2
self.name = name
def flux(self, u):
""" Numerical flux
u: conservative variables (rho, m)
F = ( m, m^2/rho + rho*a^2 )
"""
return np.array([ u[1], u[1]**2/u[0]+u[0]*self.a2])
def A(self, u):
""" Convection matrix, A; input: conservative variables """
vel = u[1]/u[0]
return np.array([[ .0, 1. ], [ -vel**2+self.a2, 2*vel]])
def s(self, u):
""" Eigenvalues of A; input: conservative variables """
vel = u[1]/u[0]
return np.array([vel-self.a, vel+self.a])
def R(self, u):
""" Right eigenvectors of A; input: conservative variables """
vel = u[1]/u[0]
return np.array([[1, vel-self.a],[1, vel+self.a]]).T
def L(self, u):
""" Left eigenvectors of A; input: conservative variables """
vel = u[1]/u[0]
return np.array([[vel+self.a, -vel+self.a],[-1., 1.]]).T / (2.*self.a)
def spectrum(self, u):
""" Spectrum of matrix A; input: conservative variables """
return self.s(u), self.R(u), self.L(u)
def roe_intermediate_state(self, u_0, u_1):
"""
rho_roe = ANY! = ... CHOOSE ONE: average = 0.5*(rho_0+rho_1)
vel_roe = (sqrt(rho_0)*vel_0+sqrt(rho_1)*vel_1)/(sqrt(rho_0)+sqrt(rho_1))
- convective/physical: (rho, vel) = ( rho_roe, vel_roe )
- conservatvie : (rho, mom) = ( rho_roe, rho_roe*vel_roe)
F_roe(u) ~ dFdu * du ~ A(u_roe(u0,u1)) * (u_1 - u_0)
"""
rho_roe = .5*(u_0[0]+u_1[0])
vel_0, vel_1 = u_0[1]/u_0[0], u_1[1]/u_1[0]
vel_roe = (vel_0*np.sqrt(u_0[0])+vel_1*np.sqrt(u_1[0])) / \
(np.sqrt(u_0[0])+np.sqrt(u_1[0]))
return np.array([rho_roe, rho_roe*vel_roe])
def absA_entropy_fix(self, u, delta=.1):
""" """
#> Eigenvalues s1 = u-a, s2 = u+a
eigvals = self.s(u)
#> Speed of sound a = .5 * ( s2 - s1 )
a = .5 * ( np.max(eigvals) - np.min(eigvals) )
#> Entropy fix
entropy_evals = np.abs(eigvals)
entropy_evals[entropy_evals < delta*a] = \
.5 * entropy_evals[entropy_evals < delta*a]**2 / ( delta * a ) + .5 * ( delta * a )
return self.R(u) @ np.diag(entropy_evals) @ self.L(u)
def roe_flux(self, u_0, u_1):
""" """
u_roe = self.roe_intermediate_state(u_0, u_1)
return .5 * (self.flux(u_1) + self.flux(u_0) + \
self.absA(u_roe) @ ( u_0 - u_1 ) )
def flux_at_boundary(self, boundary, u):
"""
boundary: boundary conditino dict
u: conservative variables (rho, m, Et) at boundary cell
"""
if ( boundary["type"] == "wall" ):
u_boundary = u.copy(); u_boundary[1] = -u[1]
if ( boundary["nor"] < 0 ):
return self.roe_flux(u_boundary, u)
else:
return self.roe_flux(u, u_boundary)
elif ( boundary["type"] == "inflow" or boundary["type"] == "outflow" ):
delta_v = self.L(u) @ ( self.cons_from_phys(boundary["phys_ext"]) - u )
eigval = self.s(u)
delta_v[ eigval*boundary["nor"] > 0 ] = .0
u_flux = u + self.R(u) @ delta_v
return self.flux(u_flux)
58.4.1.2. Numerical simulation#
58.4.1.2.1. System of equations#
Initialized the system of equations to be solved
Show code cell source
#> P-system
# Number of unknown fields, nu
# - conservative : u = (rho, mom)
# - physical/convective: p = (rho, vel), with mom = rho*vel
nu = 2
#> Parameters and initialization of the system
a = 1. # speed of sound
S = Psys1d(a)
# S = ShallowWater1d(g=1)
# S = Psys1dLinearized(a)
# S = Euler1dPIG(...)
58.4.1.2.2. Domain#
Show code cell source
#> Parameters
x0, x1 = 0., 1. # coords of left and right boundaries
ne = 200 # n. of elements
#> Domain
xs = np.linspace(x0, x1, ne+1) # coord of the cell boundaries
xc = .5 * ( xs[:-1] + xs[1:] ) # coord of the cell centers
dx = xs[1:] - xs[:-1] # volume of the cells
#> Initial condition of a Riemann problem
# ES, SS, SE, EE
rhoL, rhoR = 1., 1.
uL, uR = -.5, .5
58.4.1.2.3. Boundary conditions#
Show code cell source
# boundary_0 = { # left boundary
# "type": "wall",
# "nor" : -1.
# }
# boundary_1 = { # right boundary
# "type": "wall",
# "nor" : 1.
# }
boundary_0 = { # left boundary
"type": "inflow",
"phys_ext": np.array([rhoL, uL]), # rho_ext, u_ext
"nor": -1. # nx
}
boundary_1 = { # right boundary
"type": "outflow",
"phys_ext": np.array([rhoR, uR]), # rho_ext, u_ext
"nor": 1. # nx
}
58.4.1.2.4. Simulation parameters#
Show code cell source
#> Time
t0, t1, dt = 0., 1., .00125
nt = int((t1-t0)/dt)+1
tv = np.arange(nt)*dt
#> Boundary conditions
u0_fun = lambda t: np.array([0., 0.])
u1_fun = lambda t: np.array([0., 0.])
#> Initial boundary conditions
uc0 = np.zeros((nu,ne))
uc0[0,:] = np.where( xc<.5, rhoL, rhoR )
uc0[1,:] = np.where( xc<.5, rhoL*uL, rhoR*uR )
# uc0[0,:] = 1.
# uc0[1,:] = np.where( xc<.5, .1, -.1)
# uc0[0,:] = 1. + .2 * xc / (x1-x0)
# uc0[1,:] = 0.
58.4.1.2.5. Time loop#
Show code cell source
#> Numerical schemes
#> Time integration
time_integration = 'ee' # ...
numerical_flux = 'roe' # ...
"""
#> Boundary condiitons
u0_wall = lambda t: .0 * np.sin(2*np.pi*t) # vel at solid wall
u1_wall = lambda t: .0 * np.sin(2*np.pi*t) # vel at solid wall
r1_free = lambda t: .1 # .0 * np.sin(2*np.pi*t) # rho at free wall
"""
#> Time loop
it = 0
uuc = np.zeros((nt, nu, ne)) # Array to store solution, for small dimensional pbs
uc = uc0.copy()
uuc[0,:,:] = uc
# print(uc)
u_0, u_1 = uc[:,0].copy(), uc[:,-1].copy() # extrapolated states
while it < nt-1:
t = tv[it]
#> Evaluate fluxes at internal boundaries
if numerical_flux == 'roe':
flux = np.array([ S.roe_flux(uc[:,ia], uc[:,ia+1]) for ia in range(ne-1) ]).T
#> Flux at boundaries
flux_0 = S.roe_flux(u_0, uc[:,0])
flux_1 = S.roe_flux(uc[:,-1], u_1)
# Bcs...correction after the solution have been evaluated?
# flux = np.append(np.append(flux_0, flux), flux_1)
flux = np.concatenate((flux_0[:,np.newaxis], flux, flux_1[:, np.newaxis]), axis=1)
#> Time integration
if time_integration == 'ee':
dt = tv[it+1] - tv[it]
uc += ( flux[:,:-1] - flux[:,1:] ) * dt / dx
it += 1 # Updating counter
uuc[it,:,:] = uc.copy() # Storing results
58.4.1.2.6. Post-processing#
Show code cell source
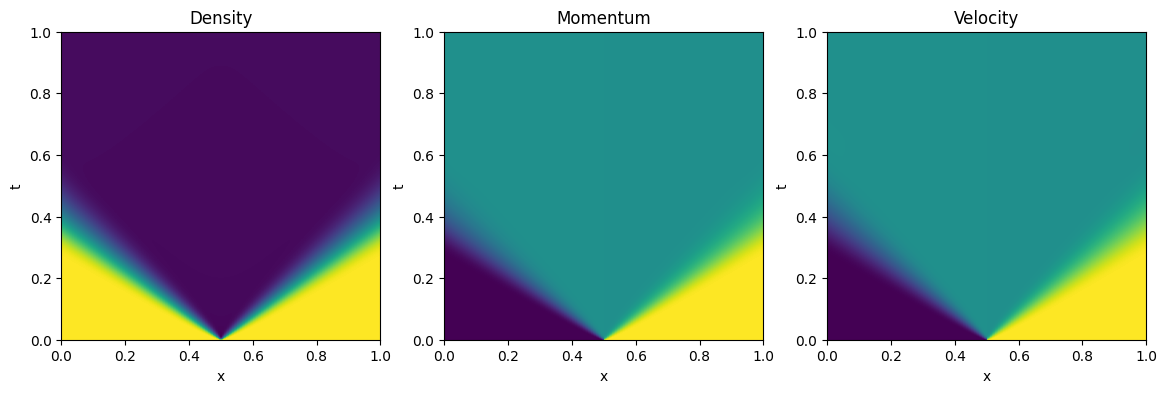
fig, ax = plt.subplots(1,3, figsize=(14, 4))
iax = 0
ax[iax].imshow(uuc[:,0,:], aspect="auto", extent=[x0, x1, t1, t0])
# ax[0].set_colorbar()
ax[iax].set_xlabel('x')
ax[iax].set_ylabel('t')
ax[iax].set_title('Density')
ax[iax].invert_yaxis()
iax = 1
ax[iax].imshow(uuc[:,1,:], aspect="auto", extent=[x0, x1, t1, t0])
# ax[0].set_colorbar()
ax[iax].set_xlabel('x')
ax[iax].set_ylabel('t')
ax[iax].set_title('Momentum')
ax[iax].invert_yaxis()
iax = 2
ax[iax].imshow(uuc[:,1,:]/uuc[:,0,:], aspect="auto", extent=[x0, x1, t1, t0])
# ax[0].set_colorbar()
ax[2].set_xlabel('x')
ax[iax].set_ylabel('t')
ax[iax].set_title('Velocity')
ax[iax].invert_yaxis()
Show code cell source
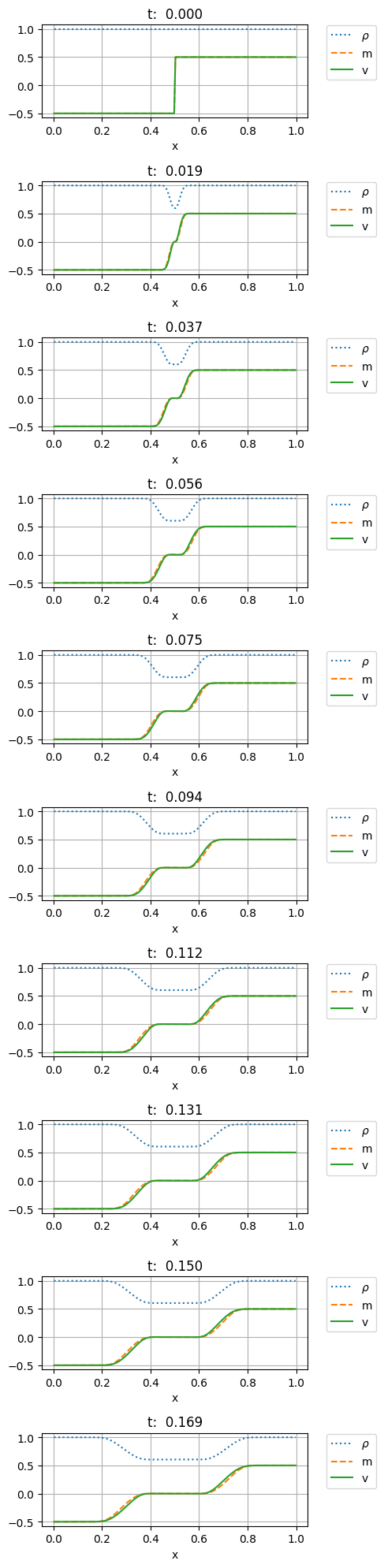
fig, ax = plt.subplots(10, figsize=(5,20))
for i in range(10):
ax[i].plot(xc,uuc[15*i,0,:], ':' , label=r'$\rho$')
ax[i].plot(xc,uuc[15*i,1,:], '--', label='m')
ax[i].plot(xc,uuc[15*i,1,:]/uuc[15*i,0,:], '-', label='v')
ax[i].set_xlabel('x')
ax[i].set_title(f"t: {15*i*dt:6.3f}")
ax[i].grid()
ax[i].legend(bbox_to_anchor=(1.05, 1.05), loc='upper left')
plt.tight_layout()